 
White data points are not significantly different. Colored and black data points are significantly over- and under-represented, respectively (adj. (B, C and E) A two-sided Fisher's exact test was employed to test for significant over- and under-representation of gene sets and P-values were adjusted for multiple testing using Bonferroni correction. The number of genes identified in each dataset as either upregulated or downregulated is compared to the sum of the remaining 19 datasets (see Supplementary Figures S3 and 4). 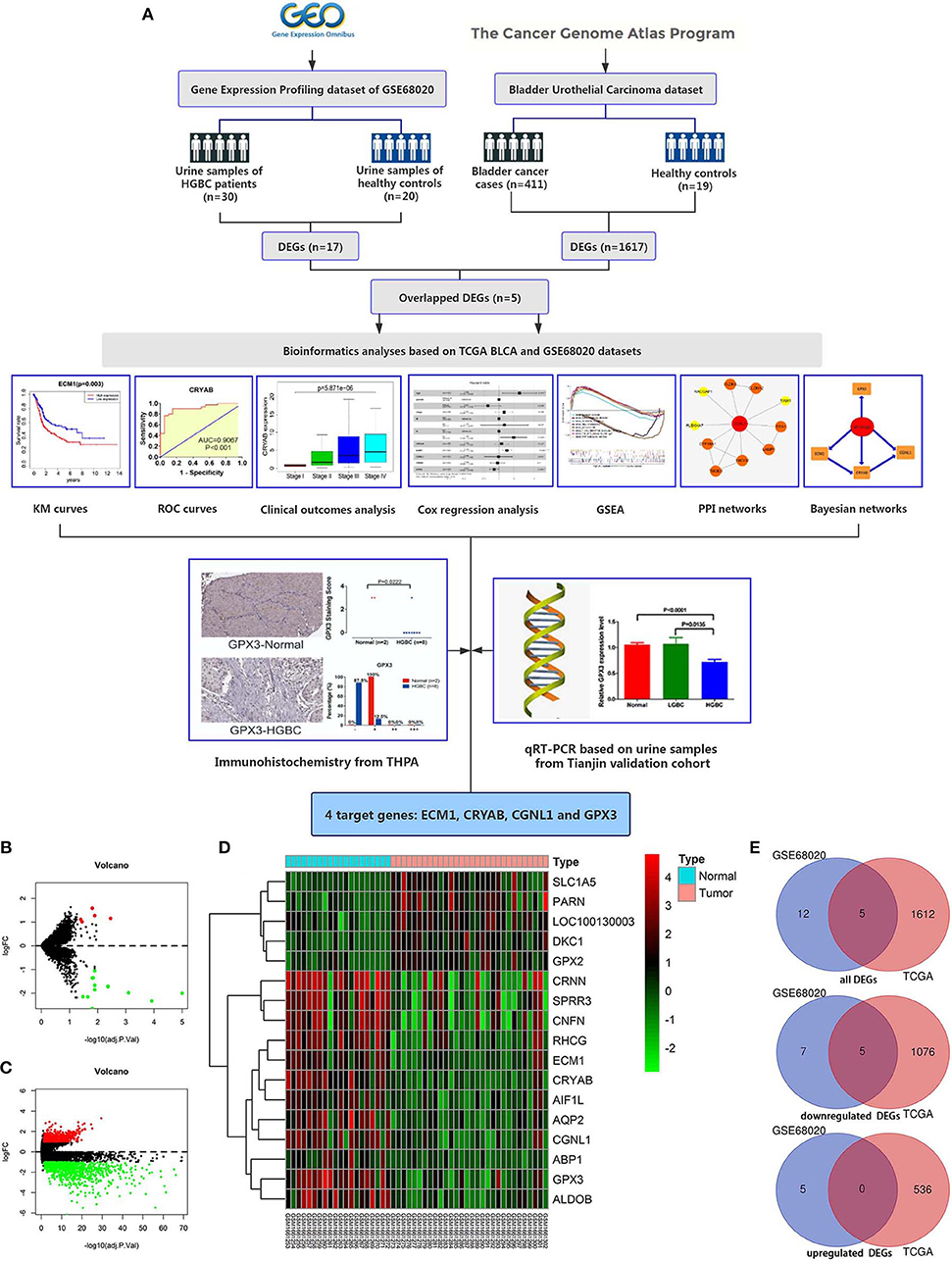
(E) Integration of 20 datasets on TP53-dependent gene regulation from multiple cell types and treatments. Correlation coefficient and two-tailed P-value was calculated using GraphPad Prism version 6.00. (C) The number of genes identified in datasets from other cell types treated with Nutlin-3a compared to the sum of the four Nutlin-3a MCF-7 datasets (see Supplementary Figure S1E–I for more) (D) Boxplot displaying the sum of the five doxorubicin datasets compared to the sum of the nine Nutlin-3a datasets. The number of genes identified in a Nutlin-3a MCF-7 dataset as either upregulated or downregulated is compared to the sum of the remaining three datasets from Nutlin-3a treated MCF-7 cells (see Supplementary Figure S1A–D for more). (B) In each dataset on TP53-dependent gene regulation, a gene can be found as upregulated ‘+1’, downregulated ‘−1’ or not regulated ‘0’. 
(A) Venn diagram displaying the overlap of genes that were detected as upregulated or downregulated by TP53 activation in datasets from Nikulenkov et al., Zaccara et al., Loayza-Puch et al. Meta-analysis of TP53-dependent gene expression. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
